10: Biohydrogen - Hydrogenases
- Page ID
- 34465
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Search Fundamentals of Biochemistry
(Learning goals written by Claude, Anthropic)
By the end of this chapter, students should be able to:
Hydrogenase Classification and [NiFe] Hydrogenases
- Classify hydrogenases into three major types ([NiFe], [FeFe], [Fe]), describe the primary physiological function of each (H₂ oxidation, H₂ production at kcat ~10,000 s⁻¹, hydride transfer), and identify the four [NiFe] subgroups (Groups 1–4) with representative physiological roles.
- Describe the subunit architecture of the membrane-bound [NiFe] MBH — large periplasmic subunit (NiFe active site), small subunit ([4Fe-3S] cluster chain anchoring to membrane), cytochrome b (quinone electron transfer) — and explain the electron flow from H₂ oxidation into the bacterial respiratory chain, connecting this to Complex II of mitochondrial electron transport.
- Explain electron bifurcation as a thermodynamic strategy — how simultaneous exergonic electron transfer to NAD⁺ (E₀' = −280 mV) subsidizes endergonic electron transfer to ferredoxin (E₀' = −500 mV) — and describe the pentameric NiFe-HydABCSL from A. mobile, identifying FMN as the bifurcation point and tracing electron flow from H₂ oxidation through FeS clusters to cofactors.
[FeFe] Hydrogenases: Structure, Mechanism, and O₂ Protection
- Describe the H-cluster active site of [FeFe]-hydrogenases — [4Fe-4S] cubane linked via Cys-503 to [2Fe₂S] subcluster with CO/CN ligands and azadithiolate (-SCH₂-NH-CH₂S-) bridge — identify the distal Fe (Fed) as the H₂ binding/production site, and explain how the hydrophobic active-site environment restricts O₂ access.
- Trace the catalytic cycle for H₂ evolution through the states Hox → Hred → HredH⁺ → HsredH⁺ → Hhyd → HhydH⁺ → Hox[H₂], identifying the oxidation states of [4Fe-4S] and [2Fe] subclusters at each step, the role of the azadithiolate nitrogen as a proton relay, and the final H₂ release from Fed.
- Describe the bifurcating [FeFe] HydABC from Thermotoga maritima as a heterododecamer, identify FMN as the bifurcation site coupling NADH oxidation to simultaneous ferredoxin reduction (endergonic) and H⁺ reduction to H₂ (exergonic), and explain the structural homology of HydA/B/C to Complex I subunits Nqo3/1/2 and its evolutionary implications.
- Describe four O₂ protection strategies in [FeFe]-hydrogenases — irreversible inactivation (no protection), CO binding forming reversible Hox-CO, sulfide binding forming Hinact via Htrans, and cysteine ligation forming Hinact by conformational change — and explain why engineering O₂ tolerance is a key challenge for biotechnological H₂ production.
Introduction
In the last section, we described different ways to produce H2 and the colors ascribed to them based on the environmental impacts. Many look to produce and use H2 to provide energy without releasing CO2. As shown in the reaction below, H2 can be used in fuel cells to power spacecraft and cars.
\begin{equation}
\begin{aligned}
& \mathrm{O}_2+4 \mathrm{H}^{+}+4 \mathrm{e}^{-} \longrightarrow 2 \mathrm{H}_2 \mathrm{O} \\
& \mathrm{H}_2 \longrightarrow 4 \mathrm{H}^{+}+4 \mathrm{e}^{-}
\end{aligned}
\end{equation}
H2-powered cars, considered a zero-emission fuel, are already available. These fully electric cars use fuel cells that oxidize H2 to generate electrical energy.
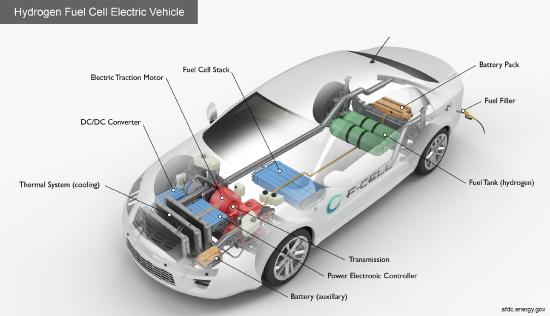
Figure \(\PageIndex{1}\): https://afdc.energy.gov/vehicles/how...tric-cars-work
Given the scale required, most H2 is produced by steam reforming of natural gas and water electrolysis. From a biochemical perspective, cells have evolved to make H2 and use H2 as an energy source. Direct microbial production of H2 is unlikely to meet society's energy needs. The 2023 International Energy Agency (IEA) report, "Hydrogen patents for a clean energy future", doesn't mention direct microbial production. However, much can be learned by studying how hydrogenases (H2ases, Hyd), enzymes that make or use H2, work. (Don't confuse hydrogenases with dehydrogenases that directly use NAD+/NADH and FAD/FADH2) for redox reactions. Transition-metal active-site mimetics can serve as potential catalysts for more industrial-scale production of H2.
Although the reversible formation of H2 involves the most elemental particles in chemistry, protons (H+) and electrons (e-), the biological reactions that produce and consume H2 are complex. Before we proceed, let's see how these reactions are similar to other biochemical reactions and pathways we have already discussed.
Use of H2 as a source of electrons for reduction reactions.
Each hydrogen in H2 has an oxidation number of 0. Each can be oxidized to H+ (oxidation number +1) with the 2 electrons transferred to a substrate/cofactor or a series of sequential substrates with higher and higher standard reduction potentials (better oxidizing agents), forming reduced products.
H2 + (substrate)OX → 2H+ + (product)RED
This general reaction is analogous to the mitochondrial electron transport chain, in which electrons are passed from a source (NADH) to oxidized acceptor molecules. The general reaction below shows each redox pair in the electron transport chain.
NADH/NAD+ → FAD/FADH2 → UQ/UQH2 → Cyto COX/Cyto CRED → O2/H2O
Some organisms have evolved to produce energy by oxidizing H2, which is analogous to the photosynthetic oxidation of water. In photosystem II, the oxygen-evolving complex oxidizes oxygen in H2O (oxidation number -2) as it forms O2 (oxidation number 0). Some redox pairs, starting with H2O/O2, are shown below for photosystem II.
H2O/O2 → P680/P680* → (Plastoquione)OX/(Plastoquione)RED
The first reaction is endergonic and requires photons as an energy source.
Use of H+ as a sink for electrons for oxidation reactions that produce H2.
H+ has an oxidation number of +1. Hence, it can be reduced to H2 (oxidation number 0) as it gains electrons from substrates/cofactors that are oxidized. This general reaction is shown below.
2H+ + (substrate/cofactor)RED → H2 + (substrate/cofactor)OX
Many microorganisms can produce H2 through variants of photosynthesis or fermentation, both of which provide the two electrons needed. E. coli has four hydrogenases (Hyd 1, 2, 3, and 4). It forms H2 through two reactions catalyzed by:
- formate (HCO2-) dehydrogenase (FDH): 2HCO2- ⇌ 2CO2 + 2H+ + 2e-
- hydrogenase (H2ase): 2H+ + 2e- → H2
The C in formate has an oxidation number of +2 and is oxidized to CO2, in which the C has an oxidation number of +4.
The formate hydrogenlyase (FHL) complex contains both the formate dehydrogenase (FDH) and a hydrogenase (H2ase) and reversibly interconverts HCO2– and H2. The E. coli FHL-1 complex, which makes H2 using fermentation, is shown below in Figure \(\PageIndex{2}\). The complex can be immobilized on a Macro-mesoporous inverse opal (IO) indium tin oxide (ITO) electrode (IO-IPO) or ITO nanoparticles (NP), which can relay electrons.
Figure \(\PageIndex{2}\): Katarzyna P. Sokol et al. J. Am. Chem. Soc. 2019, 141, 44, 17498–17502. https://doi.org/10.1021/jacs.9b09575. CC-BY license
Panel (a) shows the biological E. coli FHL-1 complex. FdhF, [Mo]-FDH; B/F/G, Fe–S cluster-containing proteins; HycE, [NiFe]-H2ase; HycD/C, membrane proteins. (17)
Panel (b) shows a IO-ITO|FDH||IO-ITO|H2ase cell: IO-ITO|FDH wired to IO-ITO|H2ase electrode.
Panel (c) shows a FDH–ITO–H2ase nanoparticle (NP) system with enzymes immobilized onto ITO NP in solution. Species size not drawn to scale.
All you need to synthesize H2 are 2 protons and 2 electrons (potentially derived from photosynthesis). Let's take a deeper look at the hydrogenase that catalyzes H2 production.
Hydrogenases (H2ases)
Hydrogenases catalyze the reversible conversion of 2H+ → H2. A hydrogenase database, HydDB, a web tool for hydrogenase classification and sequence analysis, shows their high diversity and metabolic roles. There are three classes of hydrogenases: the Ni-Fe (most abundant, primarily for H2 conversion to 2H+), the Fe-Fe (highest kcat for H2 production), and the single Fe hydrogenases, as shown in Table \(\PageIndex{2}\) below. We won't discuss the single Fe hydrogenases.
| CLASSES AND SUBCLASSES OF HYDROGENASES | ||
| [NiFe] Group 1: Respiratory H2-uptake [NiFe]-hydrogenases | ||
| 1a | Periplasmic | Electron input for sulfate, metal, and organohalide respiration. [NiFeSe] variants. |
| 1b | Prototypical | Electron input for sulfate, fumarate, metal, and nitrate respiration. |
| 1c | Hyb-type | Electron input for fumarate, nitrat,e and sulfate respiration. Physiologically reversible. |
| 1d | Oxygen-tolerant | Electron input for aerobic respiration and oxygen-tolerant anaerobic respiration. |
| 1e | Isp-type | Electron input primarily for sulfur respiration. Physiologically reversible. |
| 1f | Oxygen-protecting | Unresolved role. May liberate electrons to reduce reactive oxygen species. |
| 1g | Crenarchaeota-type | Electron input primarily for sulfur respiration. |
| 1h | Actinobacteria-type | Electron input for aerobic respiration. Scavenges electrons from atmospheric H2. |
| 1i | Coriobacteria-type (putative) | Undetermined role. May liberate electrons for anaerobic respiration. |
| 1j | Archaeoglobin-type | Electron input for sulfate respirationπ. |
| 1k | Methanophenazine-reducing | Electron input for methanogenic heterodisulfide respiration. |
| [NiFe] Group 2: Alternative and sensory uptake [NiFe]-hydrogenases | ||
| 2a | Cyanobacteria-type | Electron input for aerobic respiration. Recycles H2 produced by other cellular processes. |
| 2b | Histidine kinase-linked | H2 sensing. Activates a two-component system controlling hydrogenase expression. |
| 2c | Diguanylate cyclase-linked (putative) | Undetermined role. May sense H2 and regulate processes through cyclic di-GMP production. |
| 2d | Aquificae-type | Unresolved role. May generate reductant for carbon fixation or have a regulatory role. |
| 2e | Metallosphaera-type (putative) | Undetermined role. May liberate electrons primarily for aerobic respiration. |
| [NiFe] Group 3: Cofactor-coupled bidirectional [NiFe]-hydrogenases | ||
| 3a | F420-coupled | Couples oxidation of H2 to the reduction of F420 during methanogenesis. Physiologically reversible. [NiFeSe] variants. |
| 3b | NADP-coupled | Couples oxidation of NADPH to the evolution of H2. Physiologically reversible. May have sulfhydrogenase activity. |
| 3c | Heterodisulfide reductase-linked | Bifurcates electrons from H2 to heterodisulfide and Fdox in methanogens. [NiFeSe] variants. |
| 3d | NAD-coupled | Interconverts electrons between H2 and NAD depending on cellular redox state. |
| [NiFe] Group 4: Respiratory H2-evolving [NiFe]-hydrogenases | ||
| 4a | Formate hydrogenlyase | Couples formate oxidation to fermentative H2 evolution. May be H+-translocating. |
| 4b | Formate-respiring | Respires formate or carbon monoxide using H+ as an electron acceptor. Na+-translocating via Mrp. |
| 4c | Carbon monoxide-respiring | Respires carbon monoxide using H+ as an electron acceptor. H+-translocating. |
| 4d | Ferredoxin-coupled, Mrp-linked | Couples Fdred oxidation to H+ reduction. Na+-translocating via Mrp complex. |
| 4e | Ferredoxin-coupled, Ech-type | Couples Fdred oxidation to H+ reduction. Physiologically reversible via H+/Na+ translocation. |
| 4f | Formate-coupled (putative) | Undetermined role. May couple formate oxidation to H2 evolution and H+ translocation. |
| 4g | Ferredoxin-coupled (putative) | Undetermined role. May couple Fdred oxidation to proton reduction and H+/Na+ translocation. |
| 4h | Ferredoxin-coupled, Eha-type | Couples Fdred oxidation to H+ reduction in anaplerotic processes. H+/Na+-translocating. |
| 4i | Ferredoxin-coupled, Ehb-type | Couples Fdred oxidation to H+ reduction in anabolic processes. H+/Na+-translocating. |
| [FeFe] Hydrogenases | ||
| A1 | Prototypical | Couples ferredoxin oxidation to fermentative or photobiological H2 evolution. |
| A2 | Glutamate synthase-linked (putative) | Undetermined role. May couple H2 oxidation to NAD reduction, generating reductant for glutamate synthase. |
| A3 | Bifurcating | Reversibly bifurcates electrons from H2 to NAD and Fdox in anaerobic bacteria. |
| A4 | Formate dehydrogenase-linked | Couples formate oxidation to H2 evolution. Some bifurcate electrons from H2 to ferredoxin and NADP. |
| B | Colonic-type (putative) | Undetermined role. May couple Fdred oxidation to fermentative H2 evolution. |
| C1 | Histidine kinase-linked (putative) | Undetermined role. May sense H2 and regulate processes via histidine kinases. |
| C2 | Chemotactic (putative) | Undetermined role. May sense H2 and regulate processes via methyl-accepting chemotaxis proteins. |
| C3 | Phosphatase-linked (putative) | Undetermined role. May sense H2 and regulate processes via serine/threonine phosphatases. |
| [Fe] Hydrogenases | ||
| All | Methenyl-H4MPT dehydrogenase | Reversibly couples H2 oxidation to 5,10-methenyltetrahydromethanopterin reduction. |
Dan Søndergaard et al., Scientific Reports volume 6, Article number: 34212 (2016). Creative Commons Attribution 4.0 International License. http://creativecommons.org/licenses/by/4.0/.
The three main types have different main functions in general. The Ni-Fe, Fe-Fe, and Fe H2ases generally oxidize H2, produce H2, and promote H- (hydride) transfer, respectively, as shown in Figure \(\PageIndex{3}\) below.
Figure \(\PageIndex{3}\): The active site structures of [NiFe] H2ases that mainly catalyze H2 oxidation reactions, [FeFe] H2ases that mainly catalyze H2 evolution reactions, and [Fe] H2ases that catalyze H−transfer to the substrate via heterolytic H2 cleavage. X, possible H2 active sites; Y, methenyltetrahydromethanopterin; GMP, guanosine monophosphate. Seiji Ogo et al., Science Advances.(2020). DOI: 10.1126/sciadv.aaz81. Creative Commons Attribution-NonCommercial License 4.0 (CC BY-NC).
Much effort has been devoted to developing transition-state analogs of the active site to act as catalysts for H2 production in fuel cells. Transition-metal catalysts that mimic the structures and activities of the three hydrogenases have been developed. Figure \(\PageIndex{4}\) shows three specific ones below.
Figure \(\PageIndex{4}\): The differing reactivity of the three isomers. Y′, methylene blue [MB]+.Seiji Ogo et al., ibid.
The ligand containing P and PH is bis(diphenylphosphino)ethane.
First, we will explore the Ni-Fe hydrogenases.
Ni-Fe H2ases (Hyd):
We'll discuss two examples of Ni-Fe H2ases
Group 1a periplasmic (membrane-bound) hydrogenases - MBH
These are used in fuel cells and H2-producing devices because they can adhere to surfaces and serve as useful, heterogeneous (not in solution) catalysts. O2 also damages some. The enzyme consists of a large periplasmic subunit and a small subunit that anchors the protein to the bacterial plasma membrane. This enzyme oxidizes H2: H2 → 2H++2e−. The electrons enter the bacterial respiratory chain through quinones. The transmembrane part of the small subunit binds cytochrome b, which is involved in electron transfer with the quinones, as we saw in Complex II of mitochondrial electron transport. Some soil bacteria (like Ralstonia eutropha) can use H2 as their sole energy source. The orientation of a NiFe MBH within a bacterial cell is shown in Figure \(\PageIndex{5}\) below.
Figure \(\PageIndex{5}\): The orientation of a NiFe MBH within a bacterial cell. Lindsey A. Flanagan* and Alison Parkin. Biochem Soc Trans. 2016 Feb 15; 44(1): 315–328 (2016). doi: 10.1042/BST20150201 Creative Commons Attribution License 3.0.
Panel (A) shows a cartoon depiction of a NiFe MBH's location within the cytoplasmic membrane. White boxes represent the redox-active metal centers, and blue, orange, and purple blocks indicate the large, small, and cytochrome subunits, respectively.
Panel (B) shows how the E. coli hydrogenase-1 large (blue ribbon), small (orange ribbon), and cytochrome (purple ribbon) subunits can interact.
Figure \(\PageIndex{6}\) shows an interactive iCn3D modelof the O2-Tolerant Membrane-Bound Hydrogenase 1 from Escherichia coli in Complex with Its Cognate Cytochrome b (4GD3). The same color coding is used for the subunits as in the above figures.
Figure \(\PageIndex{6}\): O2-Tolerant Membrane-Bound Hydrogenase 1 from Escherichia coli in Complex with Its Cognate Cytochrome b (4GD3). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...9p4CSQRDPM8nL7
The large and small chains of hydrogenase are shown in blue and orange, respectively, while cytochrome b is shown in magenta. Cofactors and key metals are shown in spacefill. F4S is the Fe4-S3 cluster, SF4 is the iron-sulfur cluster, HEM is hemoglobin, FCO is carbon monoxide-(dicyano) Fe, and Ni is nickel.
The biological (functional) unit consists of two heterodimers. At low O2 levels, a single dimer can reactivate the other dimer upon exposure to O2. The enzyme is present at its highest concentration during anaerobic fermentation. Remember that E. coli is a facultative anaerobe that can shift its metabolic pathways to fit conditions. Perhaps its primary role is to reduce O2 to water and protect enzymes sensitive to it. Cytochrome b may mostly anchor the dimeric H2ase in the membrane.
A bifurcating Ni-Fe H2ase
These enzymes are more complicated. They oxidize H2 and direct the two electrons through a complex pathway that bifurcates the flow to different substrates/cofactors. They move an electron to a low-potential (i.e., not a strong oxidizing agent) high-energy species, which is reduced in an endergonic process. The other electron simultaneously moves to a high-potential (i.e., a strong oxidizing agent), low-energy species, which is also reduced in an exergonic process. The overall electron transfer is thermodynamically favorable. An example might prove helpful. NADH (E0' = -280 mV, higher potential) can reduce the protein ferredoxin (E0' = -500 mV, lower potential), which can then pass its electrons in other reactions, including the formation of H2, CH4, and NH3. ATP is not required.
Four classes of bifurcating enzymes that use FAD/FADH2 or FMN/FMNH2 are known. They are optimal because they can participate in either 1 or 2-electron transfers. We will see an example of a Fe-Fe H2ase further below.
In electron transport, we encountered an electron bifurcating complex in the Q-cycle of Complex III. Reduced ubiquinone (UQH2, or ubiquinol) is oxidized, and the two lost electrons are bifurcated to cytochrome C in a high-potential pathway and to UQ to reform UQH2, as shown in Figure \(\PageIndex{7}\) below.
Figure \(\PageIndex{7}\): Electron bifurcation in Complex III
One example of a bifurcating Ni-Fe H2ase is the NiFe-HydABCSL protein from the bacteria A. mobile. The general structure of the pentameric form of the functional decamer is shown in Figure \(\PageIndex{8}\) below.
Figure \(\PageIndex{8}\): Structure of the A. mobile NiFe-HydABCSL pentamer. XIANG FENG et al. SCIENCE ADVANCES. 2022. DOI: 10.1126/sciadv.abm7546. Creative Commons Attribution License 4.0 (CC BY). https://creativecommons.org/licenses/by/4.0/
The five subunits are called HydA (Hyd = hydrogenase), HydB, HydC, HydL (large subunit), and HydS (small subunit). Pane (A) shows the domain structure of the five subunits. The NiFe-HydB NTD and CTDs are partially flexible, as indicated by dashed outlines. Panel (B) shows the subunit organization of the NiFe-Hyd complex and their associated cofactors.
NiFe-HydABCSL hydrogenase can reversibly oxidize H2 with 2 electrons, reducing ferredoxin in an endergonic process and NAD in an exergonic process. FMN is surrounded by a FeS cluster and appears to be the bifurcation point. The reaction is as follows:
- The HydL oxidizes H2 with two electrons passing through the FeS centers in HydA to HybB.
- The electrons are passed to FMN, where the bound NAD gets reduced.
Figure \(\PageIndex{9}\) jdkfjdkjfdkjff
Figure \(\PageIndex{9}\): Proposed mechanism of electron bifurcation/confurcation in A. mobile NiFe-HydABCSL.
(A) Overall electron transfer pathway, highlighting the three branches of the electron transfer path. The mid-potential path is a black dashed line, the exergonic path is a blue dashed line, and the endergonic path is a red dashed line. (B) Conformational changes in the HydBC bifurcation core from the electron bifurcation state (BR state) to the electron transduction state (PB state).
Figure \(\PageIndex{10}\) shows an interactive iCn3D modelof the electron bifurcating Ni-Fe hydrogenase complex HydABCSL in FMN/NAD(H) bound state 7T30
Figure \(\PageIndex{10}\): Electron bifurcating Ni-Fe hydrogenase complex HydABCSL in FMN-NAD(H) bound state (7T30). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...G9nKGwqvuuh6G7
Figure \(\PageIndex{11}\) shows an interactive iCn3D modelof the cofactors in the electron bifurcating Ni-Fe hydrogenase complex HydABCSL in FMN/NAD(H) bound state 7T30
Figure \(\PageIndex{11}\): Cofactors in the electron bifurcating Ni-Fe hydrogenase complex HydABCSL in FMN-NAD(H) bound state (7T30). (Copyright; author via source). Click the image for a popup, or use this external link: https://structure.ncbi.nlm.nih.gov/i...hHuzaWUDYCMnAA.
Zoom in on the Ni-Fe catalytic site. The ligands that form coordinate covalent bonds to the Fe are called FCC, or carbon monoxide-(dicyano)-Fe, Figure \(\PageIndex{12}\) below. There are also bridging sulfurs between Fe and Ni.
Figure \(\PageIndex{12}\): Carbonmonoxo-dicyano-Fe and is shown in detail in
Fe-Fe hydrogenases
These enzymes catalyze various reactions as illustrated in Figure \(\PageIndex{13}\) below.
Figure \(\PageIndex{13}\): [FeFe]-hydrogenases phylogeny and known functions. Morra S. Front Microbiol. 2022 Mar 2;13:853626. doi: 10.3389/fmicb.2022.853626. PMID: 35308355; PMCID: PMC8924675. Creative Commons Attribution License (CC BY)
As previously proposed, a phylogenetic tree shows the evolutionary relationships among [FeFe]-hydrogenase sequences from public databases. Enzymes experimentally characterized are indicated on the tree to show their relative position. The proposed physiological function of each enzyme is also presented where known. Hyd, hydrogenase subunit; FdhF, formate dehydrogenase subunit; Fdrex/ox, reduced/oxidized ferredoxin; NADH/NAD+, reduced/oxidized nicotinamide adenine dinucleotide. They are found in prokaryotic and eukaryotic microorganisms but not in Archaea.
These are the most active for H2 production with a kcat around 10,000 s-1. They contain a Fe2S2 cluster with CO and CN ligands forming bonds to the iron, with the iron ions bridged by a -SCH2-NH-CH2S- (aza-dithiolate). A cysteine links the Fe2S2 to a Fe4S4 cluster. These two are called the H-cluster (or [Fe]H. Within this class are a soluble, monomeric cytoplasmic form, a heterodimeric periplasmic form, and a soluble, monomeric chloroplastic form. This one has a ferredoxin, connecting it to the electron transport chain in photosynthesis. Some in this group, using both NADH and ferredoxin, are called bifurcating types, as they send two electrons from a donor in two different directions. More on this later.
They contain multiple FeS clusters. The H-cluster consists of a Fe2S2 cluster linked to a Fe4S4 (cubane-like) cluster via a cysteine. The Fe2S2 group has CO and CN ligands, with the two Fe ions of the Fe2S2 unit coordinated by an azadithiolato ligand, as shown below in Figure \(\PageIndex{14}\).
Figure \(\PageIndex{14}\): Chemical structure of the H-cluster, which is the active site of the [FeFe] hydrogenase enzyme. Rakesh C. Puthenkalathil et al., Phys. Chem. Chem. Phys., 2020, 22, 10447. https://pubs.rsc.org/fa/content/arti.../cp/c9cp06770a. This article is licensed under a Creative Commons Attribution 3.0 Unported License
The Fe in the [2Fe2S] cluster is linked to the [4Fe-4S] cubane and has six ligands, so it is saturated. The other Fe has an extra coordination site denoted by X, which can bind H+ or H2. The cluster is buried in a hydrophobic catalytic site, which helps restrict O2 access.
As we did for the Fe-Ni H2ases, we will study two examples of Fe-Fe H2ases.
Fe-Fe hydrogenase (CpI) from Clostridium pasteurianum
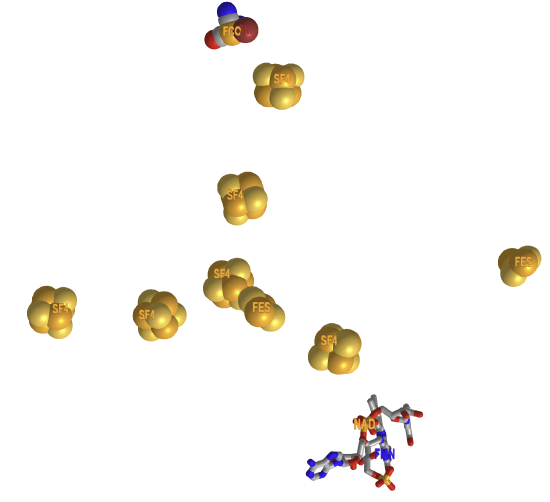
Figure \(\PageIndex{15}\) shows an interactive iCn3D modelof the H-Cluster (HC1) of Fe-Fe hydrogenase (CpI) from Clostridium pasteurianum (1FEH).
Figure \(\PageIndex{15}\): H-Cluster (HC1) of Fe-Fe hydrogenase (CpI) from Clostridium pasteurianum (1FEH). (Copyright; author via source). Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...TksW8FGn7crh39
In this model, all CN ligands are shown as CO. The sulfurs are shown in green. Hover over the atoms/ions to identify them. (In iCn3D, choose, Select, Select on 3D, atom). The [4Fe-4S] subcluster forms coordinate covalent bonds with four cysteines (300, 355, 499, and 503), with one cysteine (503) forming a bridge to the [2Fe] cluster. The Fe ions in that cluster are surrounded by an octahedral arrangement of ligands. One of the ligands is water (no connecting C atom).
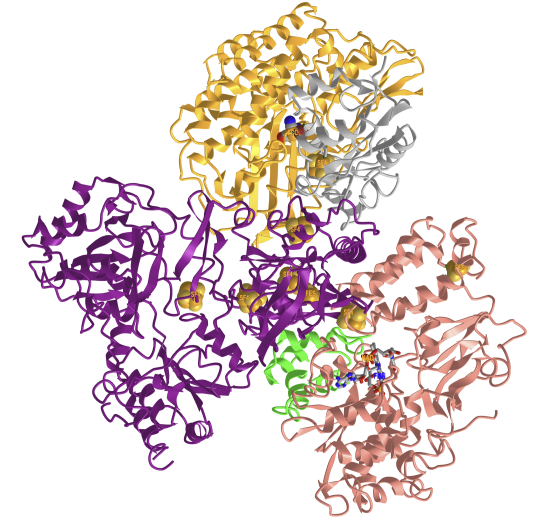
Figure \(\PageIndex{16}\) shows an interactive iCn3D modelof the Fe-Fe hydrogenase (CpI) from Clostridium pasteurianum (1FEH)
Figure \(\PageIndex{16}\): Fe-Fe hydrogenase (CpI) from Clostridium pasteurianum (1FEH). (Copyright; author via source).
Click the image for a popup or use this external link: https://structure.ncbi.nlm.nih.gov/i...QJsoZ8zbvkpDc7.
As we mentioned above, the net reaction is:
2H+ + (substrate/cofactor)RED → H2 + (substrate/cofactor)OX
Many microorganisms can produce H2 through variants of photosynthesis or fermentation, both of which provide the two electrons needed. E. Coli has four hydrogenases (Hyd 1, 2, 3, and 4). It forms H2 through two reactions catalyzed by:
- formate (HCO2-) dehydrogenase: 2HCO2- ⇌ 2CO2 + 2H+ + 2e-
- hydrogenase 3: 2H+ + 2e- → H2
Figure \(\PageIndex{17}\) below shows a reaction scheme for the production of H2 linked to photosystem I in the chloroplast of microalgae under anaerobic conditions. It starts with the absorption of a photon by P700, which, in the excited state, transfers an electron to a 4Fe4S cluster in ferredoxin. This transfers an electron to the HC-cluster, then to H+.
Figure \(\PageIndex{17}\): Schematic representation of electron flow from Photosystem I to an [FeFe]-hydrogenase via a ferredoxin redox mediator (Photosystem I). JuanAmaro-Gahete et al., Coordination Chemistry Reviews. 448, December 2021. https://doi.org/10.1016/j.ccr.2021.214172. Creative Commons CC-BY
A possible mechanism for forming H2 in the H cluster is shown below in Figure \(\PageIndex{18}\).
Figure \(\PageIndex{18}\): Proposed mechanistic cycle for hydrogen evolution in the H cluster by [FeFe]-hydrogenase adapted from Lubitz et al. JuanAmaro-Gahete et al., ibid
Start at the top left, which shows the rest of the oxidized state. In the enzyme's most oxidized resting system (Hox), the [4Fe4S] cubane is in a 2 + oxidation state while the catalytic subcluster [2Fe] is a mixed-valence FeIFeII state. The first one-electron reduction results in the formation of the Hred state, in which the [4Fe4S] subcluster is reduced to the 1+ oxidation state. Protonation of the N of aza-propane-1,3-dithiolate ligand (adt-N) triggers an intramolecular charge shift to form HredH+ in which the [4Fe4S] cubane is in the 2 + state and the [2Fe] subsite is reduced to a homovalent FeIFeI state. Subsequent one-electron reduction of the subcluster [4Fe4S] gives rise to the “super-reduced” state HsredH+. In the next step of the catalytic cycle, an intermediate hydride state [Hhyd] is formed by an intramolecular proton shift from the adt-N to the distal iron Fd. This process is coupled to an electron rearrangement in the [2Fe] subsite, leading to a formal FeIIFeII oxidation state. The addition of a second proton coupled to another charge shift from the reduced [4Fe4S] to the [2Fe] subsite, either in one or two discrete steps, gives rise to [HhydH+] that is characterized by a formal FeIFeII oxidation state. At this point, there is an equilibrium between the HhydH+ and Hox[H2] in which the hydride and the proton are combined into a hydrogen molecule at the distal iron of the system. The catalytic cycle is closed by H2 release, returning to the initial Hox configuration.
A bifurcating [Fe-Fe] hydrogenase from Thermotoga maritima (HydABC)
This enzyme, functionally a heterododecamer, uses NADH as a source of electrons, which passes them to FMN, the bifurcation site, with one electron going to oxidized ferredoxin (Fdox) and the other to H+ for reduction to FdRED and H2. The enzyme consists of a dimer of trimeric subunits HydA, HydB, and HydC, with each dimer (HydABC)2 interacting with another (HydABC)2 to form a heterododecamer; both halves acting independently. The two trimers (HydABC) in the dimer (HydABC)2 are connected by a [4Fe–4S] cluster. A flexible loop in the B and A chain has a "closed" and "open" bridge conformation, with a nearby Zn2+ important in the loop conformation.
Figure \(\PageIndex{19}\) below shows the HydABC tetramer's cryo-EM structure and the redox cofactors' arrangement.
Figure \(\PageIndex{19}\): Cryo-EM structure of the HydABC tetramer and arrangement of the redox cofactors. Chris Furlan et al. (2022) eLife 11:e79361. https://doi.org/10.7554/eLife.79361. Creative Commons Attribution License
Panel (A) shows the unsharpened 2.3 Å map of Hyd(ABC)4 with D2 symmetry enforced, showing a tetramer of HydABC heterotrimers. All four copies of HydB and C are colored blue and green, respectively. The four HydA copies that make up the complex's core are orange, yellow, pink, and red. The top and bottom halves of the complex are constituted by dimers of HydABC protomers (each HydABC unit is a protomer); the two protomers within the same dimer are strongly interacting, while a weaker interaction is present between the top and bottom dimers.
Panel (B) shows the HydABC dimer, highlighting the iron-sulfur clusters and flavin mononucleotide (FMN) constituting the electron transfer network.
Panel (C) shows the arrangement of redox cofactors within the protein complex, with two independent, identical redox networks (dashed circles); each network comprises iron-sulfur clusters belonging to a Hyd(ABC)2 unit, consisting of two strongly interacting HydABC protomers.
Panel (D) shows a schematic of the electron-transfer network for one of the two identical Hyd(ABC)2 units, with edge-to-edge distances (in Å) between the various cofactors. Note that our structure is of apo-HydABC and contains only the [4Fe–4S]H subcluster of the H-cluster. The 2H+/H2 interconversion reaction in (B) illustrates the site at which this reaction occurs, but this will only occur in the fully assembled H-cluster, including [2Fe]H.
Figure \(\PageIndex{20}\) shows an interactive iCn3D modelof the electron-bifurcating [FeFe] hydrogenase from Thermotoga maritima (HydABC) (7P5H), using the same colors as the figure above.
Figure \(\PageIndex{20}\): electron-bifurcating [FeFe] hydrogenase from Thermotoga maritima (HydABC) (7P5H). (Copyright; author via source).
Click the image for a popup, or use this external link: https://structure.ncbi.nlm.nih.gov/i...2hwCUnJ9BgwJi8.
Only the dimer (HydABC)2 is shown. The A (gold), B (blue), and C (green) chains are colored as in the previous figure. The conformationally flexible loop at the C-terminal of a B chain in the closed state is shown in red. The gate also includes the C-terminal part of the A subunit near it.
Figure \(\PageIndex{21}\) shows the closed-bridge and open-bridge conformations of HydABC (the closed loop was shown in the model above).
Figure \(\PageIndex{21}\): closed-bridge and open-bridge conformations of HydABC from Thermotoga maritime.
Panel (D) shows the HydB bridge domain in the open position and its fitted model. Panel (E) shows a Zn2+ hinge region and the two possible conformations of the HydB bridge domain, open (blue) and closed (light blue).
The similarities in cofactor arrangement in the Thermotoga maritime Hyd A, B, and C subunits compared to the Nqo1, Nqo2, and Nqo3 subunits in Complex I from Thermus thermophilus (discussed in Chapter 19.1) are shown in Figure \(\PageIndex{22}\) below.
Figure \(\PageIndex{22}\): Comparison of the HydA, B and C subunits of the electron bifurcating [FeFe] hydrogenase from Thermotoga maritima with the Nqo3, 1 and 2 subunits from respiratory complex I from Thermus thermophilus.
Panel (A) shows the subunits HydA (red), HydB (blues), and HydC (green) overlaid with, respectively, Nqo3, Nqo1, and Nqo2 (all yellow) of complex I from T. thermophilus (Gutiérrez-Fernández et al., 2020, PDB: 6ZIY).
Panel (B) compares the NADH-binding site of the Nqo1 subunit of complex I from T. thermophilus (light blue) with the flavin mononucleotide (FMN) site in HydB; the high similarity suggests that NADH binds in the proximity of FMN in HydABC, similar to complex I.
Panel (C) shows an electron transfer network in HydABC compared to complex I from T. thermophilus, with edge-to-edge distances indicated in bold. The red, blue, and green dotted lines indicate the cofactors present in the HydA (Nqo3), HydB (Nqo1), and HydC (Nqo2) subunits, respectively. Note that our structure is of the apo-HydABC and lacks the [2Fe]H subcluster of the H-cluster. The 2H+/H2 interconversion reaction in (C) illustrates the site at which this reaction occurs, but this will only happen in the fully assembled H-cluster, including [2Fe]H.
Here is a link to a video showing the conformational change observed between the ‘Bridge closed forward’ (7P8N) and ‘Open bridge’ (7PN2) classes.
In the video, the HydB C-terminal iron-sulfur cluster domain is colored blue, and the HydA C-terminal iron-sulfur cluster domain is colored orange. The zinc ion (gray sphere) and ligating residues (three cysteine ligands and one histidine) are also shown. The location of the HydA C-terminal domain when the bridge is open is unknown, so it is shown transparently in both states for reference.
The geometric separation of catalytic sites and the bifurcation mechanism prevent these thermodynamically favored reactions from happening
-
H2 production from ferredoxin oxidation (in the absence of NADH oxidation)
-
NAD+ reduction by H2 (in the absence of ferredoxin reduction)
-
ferredoxin oxidation by NAD+
Oxygen Sensitivity of Fe-Fe H2ases
We have noted that Fe-Fe H2ases can be sensitive to O2. A possible mechanism involves interacting an Fe ion (Fed, the distal Fe) with oxygen, forming damaging free radicals. As CO binds more strongly than O2 to the iron in hemoglobin, its interaction with the H-center can help protect the H2ases. Sulfides can also afford protection. These mechanisms are illustrated in Figure \(\PageIndex{23}\):
Figure \(\PageIndex{23}\): Oxygen tolerance strategies in [FeFe]-hydrogenases. Morra S, ibid.
Schematic representation of the H-cluster in the oxidized active state Hox (center). In the absence of any exogenous protectant, numerous [FeFe]-hydrogenases undergo irreversible inactivation due to H-cluster damage with loss of Fe atoms (red pathway); carbon monoxide acts as a protective agent due to its ability to form Hox-CO by binding reversibly to the H-cluster at the same site as O2 (purple pathway); in DdH, a similar mechanism occurs when sulfide binds to the H-cluster forming Hinact, via the Htrans intermediate (orange pathway); in CbA5H, a conformational change in the protein structure allows for a conserved cysteine to directly bind to the H-cluster, forming Hinact (green pathway). Fep, proximal iron atom; Fed, distal iron atom; Cys, cysteine residue.
Summary
(Summary written by Claude, Anthropic)
This chapter examines the molecular machinery of biological hydrogen metabolism — the hydrogenase enzymes — in detail, connecting their structural biochemistry to the thermodynamics of electron transfer and their relevance to biotechnological H₂ production.
Hydrogenase classification. Three structurally and functionally distinct classes of hydrogenases exist. [NiFe]-hydrogenases are the most widespread and primarily oxidize H₂, passing electrons into respiratory chains. [FeFe]-hydrogenases are the most active for H₂ production, with kcat values around 10,000 s⁻¹ — roughly 100-fold faster than [NiFe] enzymes. [Fe]-hydrogenases (single-iron, or Hmd) are found only in methanogens and catalyze hydride transfer to methenyl-tetrahydromethanopterin. Within each class, functional specialization is extensive: the [NiFe] family alone spans four major groups encompassing respiratory uptake enzymes (Group 1), sensory H₂-detecting enzymes (Group 2), cofactor-coupled bidirectional enzymes (Group 3), and respiratory H₂-evolving enzymes including formate hydrogenlyase (Group 4, subgroup 4a, as found in E. coli). Synthetic transition-metal complexes that mimic hydrogenase active sites — using ligands such as bis(diphenylphosphino)ethane — are under development as industrial H₂ production catalysts.
Membrane-bound [NiFe] hydrogenase. The prototypical Group 1 [NiFe] MBH, exemplified by E. coli hydrogenase 1, is a membrane-anchored heterotrimer. The large periplasmic subunit contains the NiFe active site where H₂ is oxidized (H₂ → 2H⁺ + 2e⁻); the small subunit harbors a chain of FeS clusters including an unusual [4Fe-3S] cluster involved in O₂ tolerance; the cytochrome b subunit anchors the complex in the membrane and transfers electrons to the quinone pool, coupling H₂ oxidation to aerobic respiration. The E. coli enzyme is most abundant during anaerobic fermentation and is proposed to protect O₂-sensitive enzymes by scavenging O₂ and reducing it to water. Soil bacteria such as Ralstonia eutropha use Group 1 hydrogenases as their sole energy source, oxidizing atmospheric H₂.
Electron bifurcation in [NiFe] hydrogenases. The bifurcating NiFe-HydABCSL from Acetomicrobium mobile illustrates a sophisticated thermodynamic strategy also seen in mitochondrial Complex III's Q-cycle. This pentameric enzyme simultaneously directs two electrons from H₂ oxidation along two paths: one electron reduces oxidized ferredoxin (Fdox, E₀' = −500 mV) in an endergonic step, while the other reduces NAD⁺ (E₀' = −280 mV) in an exergonic step. FMN, surrounded by FeS clusters in the HydB subunit, is the bifurcation point. The overall reaction is thermodynamically favorable despite the endergonic ferredoxin reduction, because the exergonic NAD⁺ reduction subsidizes it — enabling the enzyme to produce reduced ferredoxin (a potent reductant for N₂ fixation, methanogenesis, and other anabolic processes) without consuming ATP.
[FeFe]-hydrogenase active site and mechanism. The [FeFe]-hydrogenases contain the H-cluster, a uniquely complex active site: a [4Fe-4S] cubane cluster connected via a bridging cysteine to a [2Fe₂S] subcluster whose two iron atoms are coordinated by CO and CN ligands (unusual in biology) and bridged by an azadithiolate (-SCH₂-NH-CH₂S-) ligand. The distal Fe (Fed) has an open coordination site (X) for H₂ and H⁺ binding. The azadithiolate nitrogen serves as a proton relay, shuttling protons between solvent and the Fe active site. The proposed catalytic cycle begins with the oxidized resting state (Hox), where [4Fe-4S] is 2+ and [2Fe] is mixed-valence Fe(I)Fe(II). Sequential one-electron reductions and protonations cycle through Hred, HredH⁺, HsredH⁺, and Hhyd states, with a critical hydride intermediate at Fed in the Hhyd state. A second proton addition forms HhydH⁺, from which H₂ is released at Fed, regenerating Hox. In microalgal chloroplasts, electrons from PSI-excited P700 flow through ferredoxin to the H-cluster, coupling photosynthetic light reactions directly to H₂ production under anaerobic conditions.
Bifurcating [FeFe] hydrogenase from Thermotoga maritima. The HydABC enzyme from T. maritima is a heterododecamer — a dimer of tetramers, each tetramer composed of two HydABC heterotrimers — with two independent, identical redox networks. NADH is oxidized at an FMN site in HydB (structurally homologous to the FMN-binding Nqo1 subunit of mitochondrial Complex I), and electrons bifurcate: one to oxidized ferredoxin (endergonic) and one to H⁺ to produce H₂ (exergonic). A flexible loop in HydB adopts "closed" and "open" bridge conformations regulated by a nearby Zn²⁺ ion, controlling electron transfer between the two HydABC protomers. The structural homology between HydA/B/C and Complex I subunits Nqo3/1/2 — including conserved FeS cluster arrangements and FMN binding geometry — suggests a deep evolutionary connection between anaerobic H₂ metabolism and the aerobic respiratory chain, with hydrogenases potentially representing ancestors of Complex I.
Oxygen sensitivity and protection. [FeFe]-hydrogenases are generally more O₂-sensitive than [NiFe] enzymes. O₂ reaches the H-cluster through gas channels in the protein and binds at Fed, generating reactive oxygen species that destroy the [2Fe₂S] cluster irreversibly. Four protective strategies have been identified: CO binding at Fed forms a stable, reversible Hox-CO inhibited state; sulfide binding in D. desulfuricans hydrogenase forms a reversible inactive state (Hinact) via the Htrans intermediate; conformational change in CbA5H allows a conserved cysteine to directly ligate the H-cluster forming Hinact; and some enzymes have no protection and undergo irreversible inactivation. Engineering these protective mechanisms into highly active [FeFe]-hydrogenases is a major goal for biotechnological H₂ production in aerobic environments.


.png?revision=1&size=bestfit&width=434&height=435)
_from_Clostridium_pasteurianum_(1FEH).png?revision=1&size=bestfit&width=322&height=357)
_from_Clostridium_pasteurianum.png?revision=1&size=bestfit&width=348&height=466)
_(7P5H).png?revision=1&size=bestfit&width=590&height=422)